Projects
NUPACK
Molecular Programming in the Cloud
PI: Niles A. Pierce (BBE and EAS Divisions)
SASE: Cody Newman, Scholar
Over the coming decades, the fields of molecular programming, nucleic acid nanotechnology, and synthetic biology are poised to generate transformative programmable molecular and cellular technologies addressing challenges to science and society ranging from neuroscience and development to diagnosis and treatment, and from renewable energy to sustainable manufacturing. In a multi-decade effort to support these emerging fields, the Pierce lab developed NUPACK (Nucleic Acid Package), a growing software suite for the analysis and design of nucleic acid structures, devices, and systems. NUPACK algorithms are unique in treating complex and test tube ensembles containing arbitrary numbers of interacting strand species, providing crucial tools for capturing concentration effects essential to analyzing and designing the intermolecular interactions that are a hallmark of these new fields. Launched in 2007, the NUPACK web application (http://www.nupack.org/) has grown to have a major impact on nucleic acid analysis and design activities worldwide; in 2019 alone, the NUPACK web application hosted 110,000 user sessions totaling 1,500,000 screen minutes and 2,500,000 page views. NUPACK papers have been cited 2200 times.
Usage has increased to the point where NUPACK’s original 240-core compute cluster was frequently overwhelmed by the research community. The Schmidt Academy collaborated with the Pierce lab to re-architect the NUPACK web application to enable the use of a scalable cloud-based computer cluster that can resize dynamically with demand. Incorporating modern software tools and practices such as a React-Redux web application, a Kotlin/PostgreSQL driven API, and a scalable AWS Kubernetes compute cluster, the new NUPACK web application aims to enable calculations far beyond previous capabilities both in terms of scale and scientific scope. The new web application also benefits from a complete rewrite of the NUPACK scientific code base (NUPACK 4.0), introducing enhanced physical models (coaxial and dangle stacking sub-ensembles), dramatic speedups (e.g., 20-120x for test tube analysis), and increased scalability. NUPACK 4 algorithms can also be run locally using the all-new NUPACK Python module (ReadTheDocs).
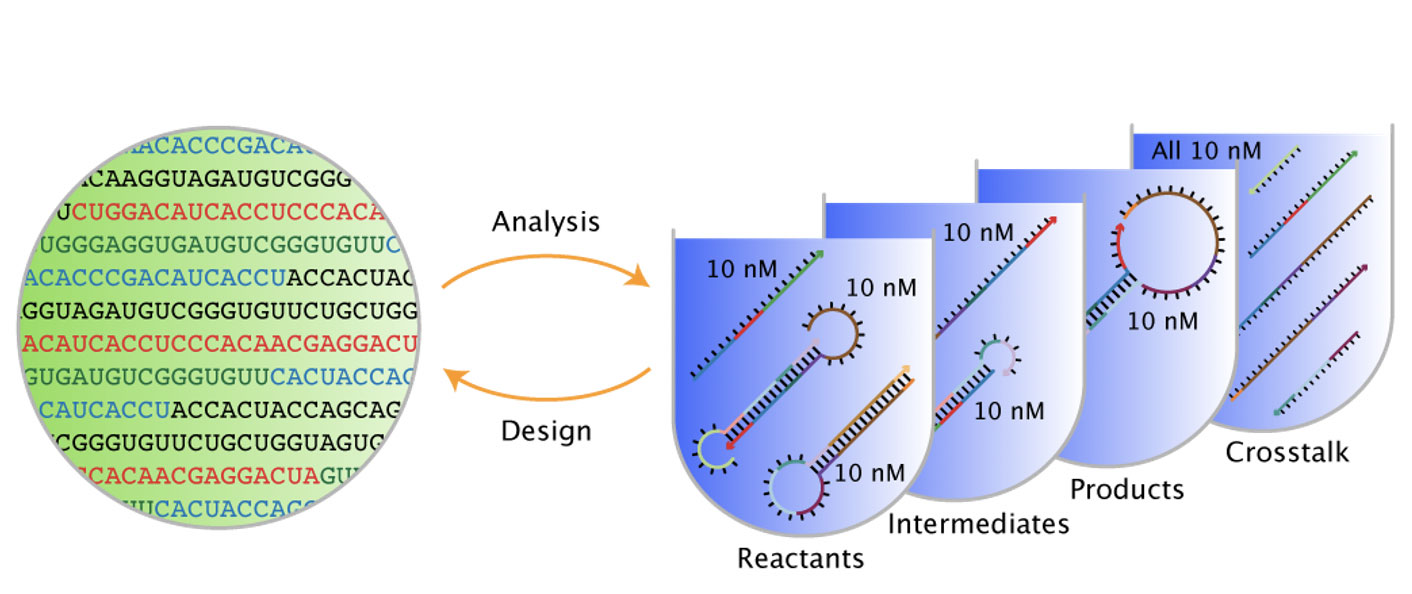
From NUPACK’s Analysis page users can analyze the equilibrium base-pairing properties one or more test tube ensembles (and subsidiary complex ensembles). From the Design page, users can design the sequences for one or more test tube ensembles (and subsidiary complex ensembles).
